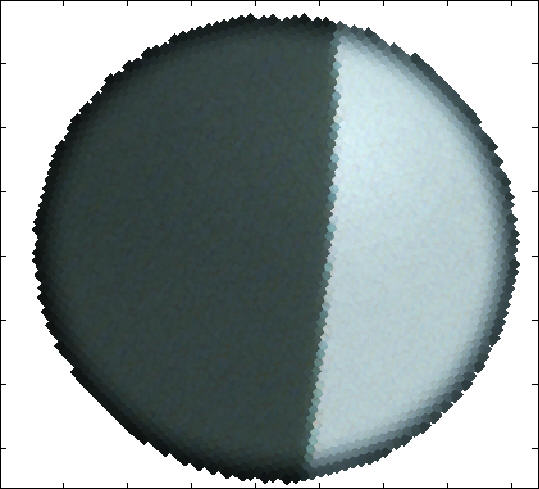
Maskfill removes features that interfere with measurements
Imatest Maskfill (mask and fill) is a standalone program for removing features in test images that interfere with Imatest analysis. A typical image is shown below: it has passed through a fiber optic bundle that adds a "honeycomb" interfering pattern. Imatest modules such as SFR will often fail with this pattern. Maskfill removes the honeycomb pattern so that Imatest modules can run properly and produce correct results.
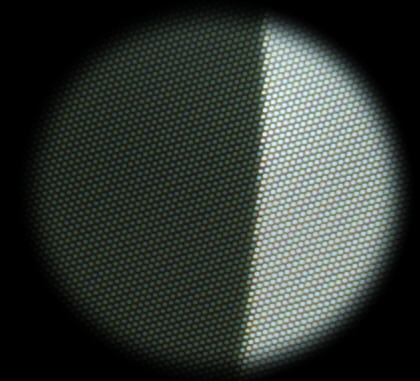

SFR slanted-edge test image with interference:
used as the Uncorrected image
Please contact us if you are interested in purchasing Maskfill.
To run Maskfill, you’ll need an image of a uniform, featureless surface (white or gray preferred) taken with the same interfering pattern as the test image(s). This pattern, called the Mask input, is used for creating a mask (the first step in Maskfill).
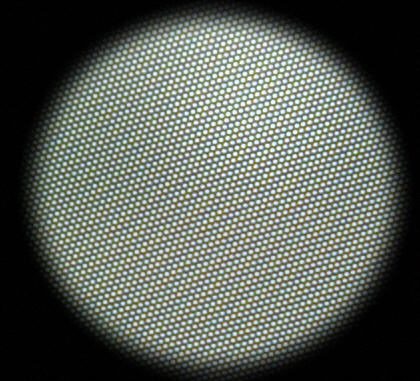

Image of plain surface:
used as the Mask input.
Must be the same pixel size and alignment as the uncorrected image.
Instructions
- Install Maskfill by double-clicking on the install file, which has a name like Maskfill-1.n.exe (where n is a number). Imatest Master should be installed first. Maskfill is installed into the Imatest program file (C:program filesImatest in English language installations).
- Acquire one or more test target images and one image of a plain, unstructured surface (the mask input). The image of the plain surface is used for making the mask: it should have the same interfering pattern as the test images. Save the images in standard file formats (TIFF, PNG, JPEG (high quality), etc.). The mask input image and the test target images must have the same alignment and the same pixel size.
- Open Maskfill by clicking on the icon on the Windows wallpaper (background) that is installed with the program, or by opening Windows Explorer and clicking on
 Maskfill in the Imatest program file. The following screen (shown reduced to 75% of the original size) appears. This window can be maximized to enhance the resolution of the images.
Maskfill in the Imatest program file. The following screen (shown reduced to 75% of the original size) appears. This window can be maximized to enhance the resolution of the images.
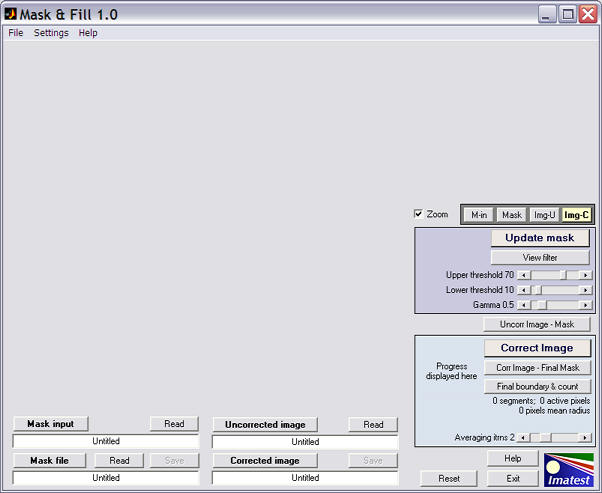
Maskfill: initial display
- The Maskfill window. Most of the left side is reserved for the main image. The lower-left contains the Display/Read/Save area for the four images processed by Maskfill. The four file names (the long white boxes) are all Untitled when Maskfill is started.
- The Mask input: the image of a plain, unstuctured surface used to create the mask: the second of the images above.
- The Mask, which can be read in or created using controls in the blue area on the right. Shown below.
- The Uncorrected image, i.e., with interference: The first of the images above.
- The Corrected image, i.e., with interference removed. Shown below.
The buttons above all files but open dialog boxes for reading the files, which will be displayed after they are read. The buttons above and save the files calculated by Maskfill. The file names replace "Untitled" in the long white boxes. The buttons on the upper right ( , , )can be used to display one of these four images on the upper right.
The large buttons above all four file boxes display the files. If or files have not been read in, the dialog box for reading them will be displayed. If hasn’t been calculated or read, it will be calculated if the file has been read in. Here is the display after Mask input (the image of the plain, unstructured surface) has been read. The long file name box for the displayed image is highlighted in yellow.
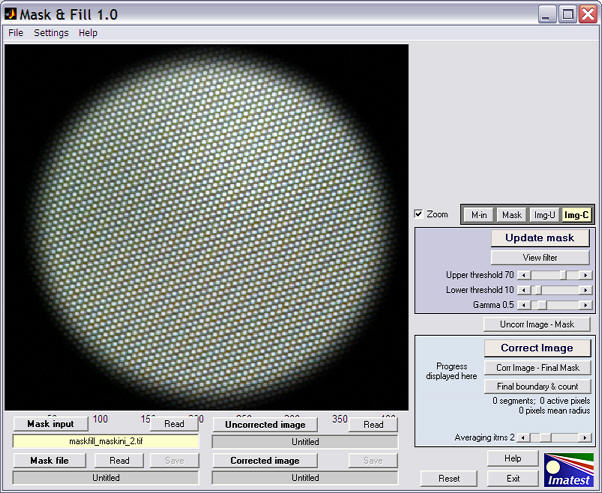
Display of Mask input image
- Read the Mask input image. This is the image of a plain, unstuctured surface used to create the mask.
- Create the mask. The light blue box to the right of the image contains the controls for creating the mask. Three sliders control the appearance of the mask.
- Upper threshold (default 70) is a variable threshold, that changes with the average (lowpass-filtered) image brightness. Increasing it shrinks the light areas of the mask.
- Lower threshold (default 10) is an absolute threshold, used so that noise in dark areas is not included in the mask.
- Gamma (default 0.5) is the encoding gamma used for creating the Mask input image file. It should be 0.5 for most standard image files (with color spaces such as sRGB, etc.) designed for display at gamma = 1.8 or 2.2. (It doesn’t have to be precise.) It should be set to 1 for linear files, i.e., files created without gamma-encoding. It affects the uniformity of the mask.
When you’ve adjusted these sliders, click to create the mask, which will appear in the main window. All desired areas should be a part of the mask (i.e., white in the B&W mask image). In this case, we want each fiber to be distinct: we don’t want them to be merged. Since the mask calculation is very fast we recommend tryuing out the slider settings to see what they do.
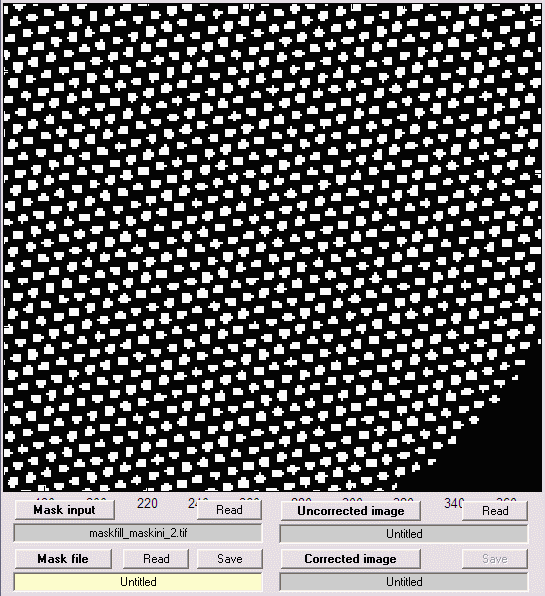
Displays can be zoomed in or out if the Zoom box is checked. You can zoom in by clicking on the image or drawing a box with the mouse. You zoom out by double-clicking on the image. A zoomed portion of the mask is shown below.
Mask image, zoomed
The button displays the lowpass-filtered version of the Mask input image, used for the Upper (variable) threshold.
- Read the Uncorrected image. This is the image that must be corrected before it can be entered into Imatest. The first image on this page is an example.
- Correct the image. The cyan box on the lower right contains the correction controls. The only adjustment is the slider that sets the maximum averaging iterations (the number of passes where adjacent pixels inside the mask (white regions) are averaged). For this image there was little improvement for more than two passes.
Press the button to perform the correction. The correction iteratively grows the image in the masked areas (white in the mask image) into adjacent unmasked areas (black in the mask image) until convergence is detected. It can be somewhat slow, but it may be sped up in future releases. The corrected image appears in the window.
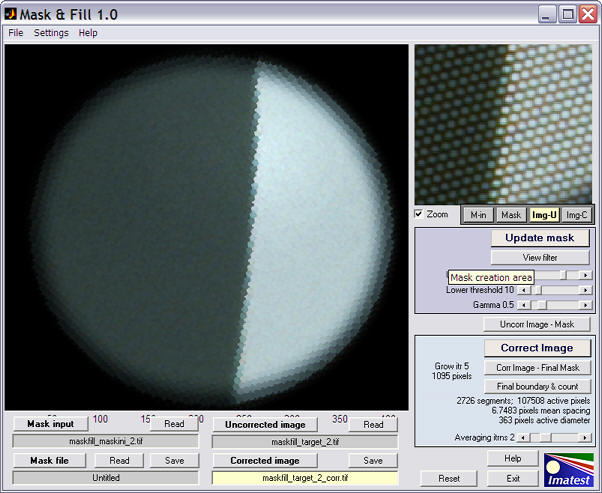
A zoomed view of the uncorrected image has been displayed in the upper-right for comparision. The Maskfill window also displays several interesting results: the number of segments (i.e., distince fiber light sources), the number of active pixels (inside the final mask), the mean spacing between the pixels (valid only for the hexagonal "honeycomb" pattern), and the diameter of the circle (4 pixels less than the measured mean of ymax-ymin and xmax-xmin for the final image boundaries).
The two buttons below can be used to produce interesting displays. The first, , shows only the active portion of the final corrected image, with the unmasked part as white. It is useful for visualizing the final mask boundary.
corrected image, unmasked area in white
The second button, , shows the final boundary and the image used to count the number of segments (light sources, i.e., active fibers in the fiber optic bundle). By zooming in (or maximizing the window), you can check to see that no individual light sources were missed or no two sources were merged. The following shows a zoom of the image. The squares represent single pixels. They are uneven in size because the Matlab display is not anti-aliased.

zoomed in, showing individual light sources
- Save the corrected image by pressing the button for the Corrected image. TIFF (uncompressed), PNG (losselessly compressed), or high quality JPEG are the preferred image formats. The image in the illustration is suitable for analysis by Imatest SFR.
This image runs nicely in Imatest SFR. Here is a sample.
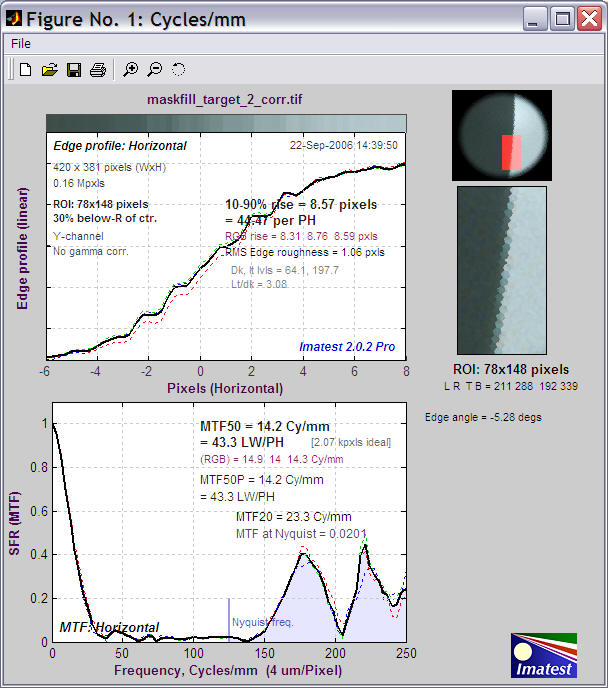
Imatest SFR results for image processed by Maskfill.